Anisotropic orientation of molecules during the deposition process can lead to built-in potentials, so called giant surface potentials, GSP, in organic thin films. As a GSP modifies the voltage drop along a device, these potentials can have a strong impact on charge carrier balance and device performance. GSP can occur in thin films of molecules with a static dipole, e.g. Alq3, but also if molecules are polarized in the thin film after deposition.
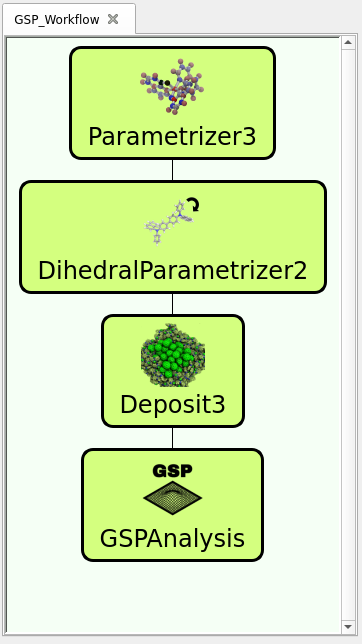
Experimentally, GSP are analyzed by measuring the potential drop of a thin film of given thickness. In our equivalent virtual experiment, we compute the GSP on digital twins of thin films by directly calculating the potential generated by the electronic structure of each molecule along the deposition axis of the thin film. In this use-case, we show you how to generate thin film morphologies of the materials Alq3 and CBP using the modules Parametrizer, DihedralParametrizer, Deposit (see details here (link to morphology generation page)), and subsequently applying the GSP Analysis tool. The workflow as predefined in the trial version of our virtual lab (available here) is illustrated in the figure to the right.
This is how you set up the simulation workflow for the GSP analysis:
- Generating an optimized 3D structure and partial charges of your input molecule, either Alq3 or CBP, using the Parametrizer. As input for the Parametrizer you can use a pre-optimized 3D structure either generated with our “synthesis” equivalent tool in the mol2 formt, or use the input provided in the trial version. The Parametrizer generates two relevant files: molecule.pdb and molecule.spf
- Load molecule.pdb and molecule.spf in the DihedralParametrizer module to generate customized energy profiles for rotation of flexible side groups of your molecule. Note that for rather rigid molecules such as Alq3, this step is not required. The relevant output of the DihedralParametrizer module is a file named dihedral_forcefield.spf
- Now use Deposit to generate a thin film of 1200 molecules. To get useful results, it is recommended to deposit a larger high but therefore narrow structure, as the GPS is measured along the deposition axis. We therefore set Lx=40, Ly=40, Lz=120 (in Angstrom). Note that these parameters define half the box size each, so the resulting morphology will extend from -40A to +40A in x and y-direction. As input for Deposit (Molecules tab), use molecule.pdb from the Parametrizer module, and (i) for Alq3, the file molecule.spf from the Parametrizer module and (ii) for CBP, the dihedral_forcefield.spf from the DihedralParametrizer module.
- In the GSPAnalysis module, load the structure.cml (i.e. the deposited morphology) from Deposit, and the molecule.spf file from the Parametrizer module. As summation method, keep the Ewald Sum option and enter the x-/y-box size (in our case: 80) in the BoxSize field.
Details on the morphology generation are available here. Note that you do not need to wait for each simulation step to finish for file transfer to the next module: You can define the data-transfer between modules using the dropdown menues next to the respective file-input fields before submitting the full workflow. See the predefined workflows in the trial or this webinar for reference.
A note on resources: Parametrizer, DihedralParametrizer and Deposit scale well for up to 32 cores. We recommend to allocate 32 cores, if available, using the Resources Tab in the GUI. The GSP analysis can be done on a single core.
Once the workflow is defined as described above, connect to your computational resource using the “Connect” button (top right in the GUI) and press Ctrl+R or “Run->run” in SimStack. Now, all you need to do is to wait for the simulation to finish.
Inspecting the results:
The GSPAnalysis will deliver you a plot, named GSP_analysis_system_scatter_ewald_slope.png , which illustrates the voltage drop directly, as visualized in the Figure below (GSPAnalysis for pristine Alq3 morphology). The experimental measurements GSP values for Alq3 and CBP of Ref 1 correspond with the calculated ones, using our method, as presented in the following table. In Ref 2 the same methodology was applied to study the GSP for a larger set of molecules, reproducing also the experimental measurements quite well.

| GSP | Experimental GSP slope / mV/nm | Calculated GSP slope mV/nm |
| Alc3 | 34-48 | 74.67 |
| CBP | 0.7 – 6.5 | 0.15 |
[1] Y. Noguchi at al., Appl. Phys.2012,111,No. 114508.
[2] P. Friederich et al. ACS Appl. Mater. Interfaces 2018, 10, 1881-1887